Program 1
Gene editing and unbiased screening for immunotherapy target discovery
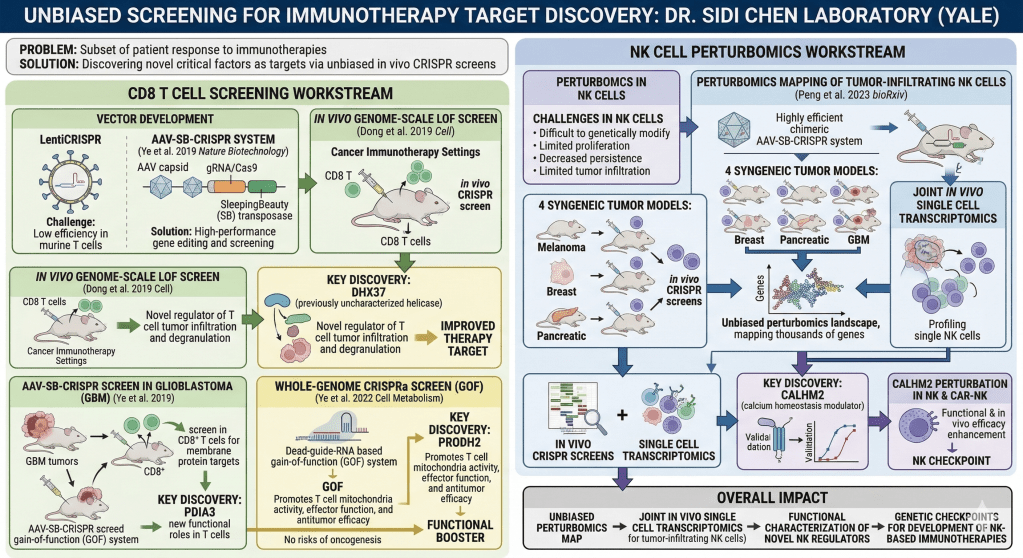
This series of work develop and apply gene editing and high-throughput screening in primary immune cells for the discovery of novel regulators as immunotherapy targets. Immunotherapy has been strikingly effective across many cancer types. However, only a subset of the patients responds to immunotherapies, urging for the discovery of critical factors as new therapeutic targets.
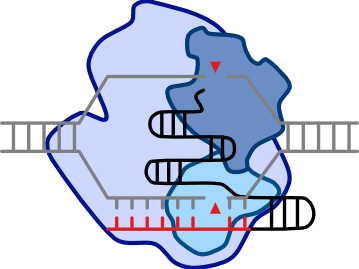
To achieve unbiased and systematic target discovery, we developed in vivo screening systems in T cells using lentiCRISPR and AAV-Transposon-CRISPR hybrid vectors. We performed the first in vivo genome-scale CRISPR screens in CD8 T cells directly under cancer immunotherapy settings and identified novel regulators of T cell tumor infiltration and degranulation, such as a previously uncharacterized helicase, DHX37 (Dong et al. 2019 Cell). As lentiCRISPR faces challenges in gene editing efficiency against murine T cells, we developed a novel and potent AAV-SleepingBeauty(SB)-CRISPR system that enables high-performance gene editing and screening in primary murine T cells (Ye et al. 2019 Nature Biotechnology). We performed an in vivo AAV-SB-CRISPR screen for membrane protein targets in CD8+ T cells in mouse models of glioblastoma (GBM), and discovered PDIA3’s new functional roles in T cells (Ye et al. 2019). By developing a dead-guide-RNA based gain-of-function (GOF) system, we achieved a whole-genome CRISPRa screen in CD8 T cell and identified functional boosters (Ye et al. 2022 Cell Metabolism). We discovered that GOF of PRODH2, a key proline metabolism enzyme, promotes T cell mitochondria activity, effector function, and antitumor efficacy, without the risks of being oncogenic (Ye et al. 2022).
Natural killer (NK) cell is an innate immune cell type that provides first level of defense. Chimeric antigen receptor (CAR) NK cells have advantages over CAR-T cells and promising therapeutic potential. Current CAR-NKs face multiple obstacles, such as lower proliferative capacity, decreased effectiveness over time, limited persistence or tumor infiltration. However, to date, there is no unbiased genetic perturbation study in primary NK cells. The challenge is that primary NK cells are difficult to genetically modify, hindering the progress of genetic studies.
To unbiasedly reveal the global genetic landscape underlying critical NK cell function, we perform perturbomics mapping of tumor infiltrating NK cells by in vivo CRISPR screens (Peng et al. 2024 Nature Biotechnology). We overcame the challenges by harnessing our highly efficient chimeric AAV-SB-CRISPR system, which enables bona fide, high efficiency gene editing in NK cells. We conducted in vivo CRISPR screens in 4 distinct, fully immunocompetent syngeneic tumor models: melanoma, breast cancer, pancreatic cancer and GBM. These screen data provided an unbiased perturbomics landscape that quantitatively mapped thousands of genes in NK cells infiltrating tumors. In parallel, we performed a joint in vivo single cell transcriptomics analysis that orthogonally identified NK cell checkpoints, by profiling tumor-infiltrating single NK cells. We discovered that CALHM2, a calcium homeostasis modulator that emerged from both screen and single cell analyses, showed both functional and in vivo efficacy enhancement when perturbed in primary NK and CAR-NK cells.
To identify gain-of-function targets that enhance CAR-NK cell efficacy, we performed an unbiased in vivo CRISPR activation screen followed by a barcoded targeted in vivo open reading frame screen in primary human CAR-NK cells. We identified and comprehensively validated OR7A10, a G protein-coupled receptor (GPCR), as the top candidate. Engineering CAR-NK cells with OR7A10 cDNA (a CRISPR-independent method with a simple manufacturing strategy) enhanced their proliferation, activation, degranulation, cytokine production, death ligand expression, chemokine receptor expression, cytotoxicity, persistence, metabolic fitness and tumour microenvironment resistance. Moreover, exhaustion in primary human NK cells derived from multiple peripheral blood and cord blood donors was reduced. OR7A10 gain-of-function CAR-NK cells displayed strong in vivo efficacy across multiple solid tumour models. For example, 100% complete response with long-term tumour control and survival benefit in an orthotopic breast cancer mouse model were achieved. These findings establish OR7A10-engineered CAR-NK cells as a highly potent and scalable off-the-shelf therapeutic for solid tumours (Yang et al. 2026 Nature).
These lines of work provided for the first time an unbiased map of perturbomics plus single cell transcriptomics for tumor-infiltrating primary NK cells and functional characterization of novel NK regulators, offering genetic checkpoints for development of NK-based immunotherapies.
Publications
Dong M*, Wang G*, Chow RD*, Ye L*, Zhu L, Dai X, Park J, Kim H, Errami Y, Guzman C, Zhou X, Chen K, Renauer P, Du Y, Shen J, Lam S, Zhou J, Lannin DR, Herbst RS and Chen S†.
Systematic identification of immunotherapy targets using genome-scale CRISPR screens in cytotoxic CD8 T cells in vivo.
Cell (2019) Aug 22;178(5):1189-1204.e23. doi: 10.1016/j.cell.2019.07.044.
https://linkinghub.elsevier.com/retrieve/pii/S0092-8674(19)30844-X
Lei Peng, Paul A Renauer, Lupeng Ye, Luojia Yang, Jonathan J Park, Ryan D Chow, Yueqi Zhang, Qianqian Lin, Meizhu Bai, Angelica Sanchez, Yongzhan Zhang, Stanley Z Lam, Sidi Chen †
Perturbomics of tumor-infiltrating NK cells.
bioRxiv 2023
doi: https://doi.org/10.1101/2023.03.14.532653
https://www.biorxiv.org/content/10.1101/2023.03.14.532653v1
Lei Peng, Paul A. Renauer, Giacomo Sferruzza, Luojia Yang, Yongji Zou, Zhenghao Fang, Jonathan J. Park, Ryan D. Chow, Yueqi Zhang, Qianqian Lin, Meizhu Bai, Angelica Sanchez, Yongzhan Zhang, Stanley Z. Lam, Lupeng Ye† & Sidi Chen†. In vivo AAV–SB-CRISPR screens of tumor-infiltrating primary NK cells identify genetic checkpoints of CAR-NK therapy.
Nature Biotechnology, 2024.
https://www.nature.com/articles/s41587-024-02282-4
Ye L*, Park JJ*, Yang Q*, Peng L*, Dong MB, Guo J, Tang E, Zhang Y, Wang G, Dai X, Du Y, Kim H, Lam S, Errami Y, Clark P, Chow RD, Montgomery R, and Chen S†.
A genome-scale gain-of-function CRISPR screen in CD8 T cells identifies proline metabolism as a means to enhance CAR-T therapy
Cell Metabolism. 2022 Apr 5;34(4):595-614.e14. doi: 10.1016/j.cmet.2022.02.009. Epub 2022 Mar 10. PMID: 35276062 PMCID: PMC8986623 (available on 2023-04-05)
https://www.sciencedirect.com/science/article/pii/S1550413122000535
Ye L*, Park JJ*, Dong MB*, Yang Q, Chow RD, Peng L, Guo J, Dai X, Wang G, Errami Y, and Chen S†.
In vivo CRISPR screening in CD8 T cells with AAV–Sleeping Beauty hybrid vectors identifies membrane targets for improving immunotherapy for glioblastoma.
Nature Biotechnology 37, 1302–1313 (2019). Sep 23. doi: 10.1038/s41587-019-0246-4. PMID: 31548728
https://www.nature.com/articles/s41587-019-0246-4
Dong MB, Tang K, Zhou X, Zhou JJ, Sidi Chen. Tumor immunology CRISPR screening: present, past and future. (2021). Trends in Cancer. DOI: doi.org/10.1016/j.trecan.2021.11.009
https://linkinghub.elsevier.com/retrieve/pii/S2405-8033(21)00239-9
Luojia Yang, Paul A. Renauer, Kaiyuan Tang, Josh Saskin, Liqun Zhou, Charles Zou, Seok-Hoon Lee, Madison Fox, Samuel Johnson-Noya, Benedict Weiss, Stephanie Deng, Paris Fang, Binfan Chen, Giacomo Sferruzza, Saba Fooladi, Kai Zhao, Daniel Park, Feifei Zhang, Jiayi Tu, Jing Chen, Jennifer Moliterno, Murat Gunel, Lei Peng† & Sidi Chen†. OR7A10 GPCR engineering boosts CAR-NK therapy against solid tumours.
Nature, 2026
https://www.nature.com/articles/s41586-026-10149-8
Program 2
Massively parallel gene editing of therapeutic immune cells to innovate next-gen cell therapies
This series of work develop massively parallel gene editing tools and apply them in immune cells to innovate next-generation cell therapies.
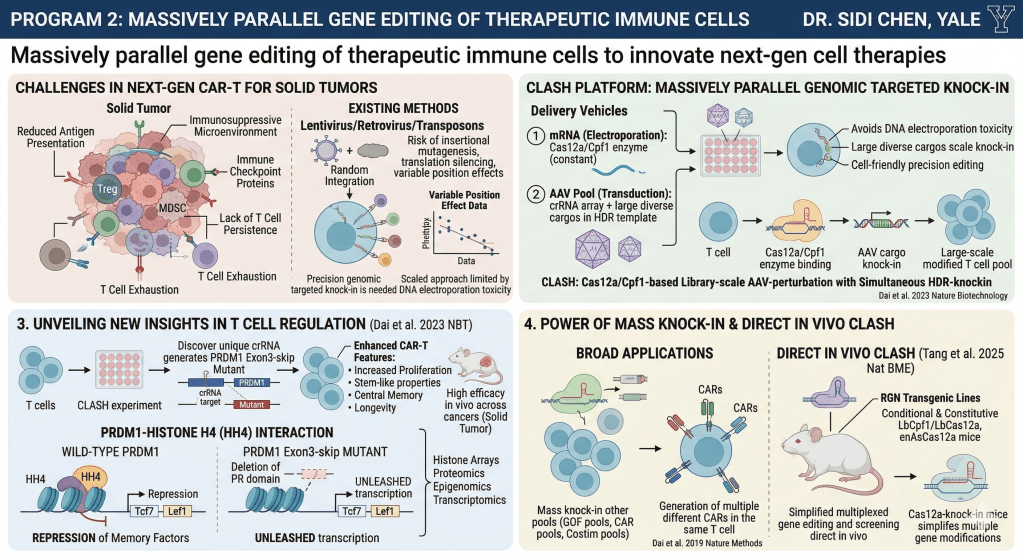
Cell therapy is a transformative class of therapeutics. However, currently approved CAR-T cell therapy faces many hurdles especially in solid tumors, where the T cell-mediated cytotoxicity against cancer can be abolished by multiple cancer-immune mechanisms, such as reduced or lost antigen presentation, immune-suppressive tumor environment, immune checkpoint proteins, lack of T cell persistence, and T cell exhaustion. Engineering more sophisticated CAR-T cells with precision control and other desired features requires a highly efficient platform.
The field has been doing genetic screens to discover new genes, however, such studies are predominantly based on lentivirus or retrovirus or transposons to integrate the target DNA sequence and, therefore, carry the risk of insertional mutagenesis and translational silencing and the possibility of variable position effects during high throughput screening. Our data showed that position effect is significant, where insertions of the same gene in two different loci can have drastically different phenotypic outcome. Genomic targeted knock-in is needed to precisely control such position effects. Cas9 RNP plus donor DNA approach has been used for knock-ins, and scaled to minipools of dozens of cargos. However, the cellular toxicity associated with DNA electroporation affects the efficiency and yield of multiplex gene modifications and naturally limits the scale of knock-in in human primary T cells. Thus, we develop a more efficient, flexible and cell-friendly precision gene editing system that enables large-scale knock-ins.
We adopted two delivery vehicles in combination: we used mRNA to deliver the gene editing enzyme Cas12a/Cpf1 via electroporation, and use AAV to deliver crRNA array and the knock-in transgene cargos embedded inside the HDR template. The mRNA component encoding the endonuclease is constant, and the AAV vector can be easily designed and scaled up to carry large diverse cargos, then synthesized in silico and packaged in vitro. With Cpf1 mRNA + AAV pool, we developed CLASH (Cas12a/Cpf1-based Library-scale AAV-perturbation with Simultaneous HDR-knockin), a novel platform for advanced massively parallel CAR-T engineering (Dai et al. 2023 Nature Biotechnology). CLASH for the first time enables massively parallel knock-in engineering in human cells. Because one of the most efficient ways to engineer superior therapeutic candidate cells is to engineer many different variants of candidates and select the best ones based on functional characteristics. CLASH enables high-throughput engineering of CAR-Ts and simultaneously identifies the best candidates for potential therapeutic applications by probing many possible candidates for their ability to enhance desirable CAR-T features.
We performed multiple CLASH experiments, in CD4, CD8 and CD3 T cells, both in vitro and in vivo. Our CLASH experiments discovered a unique crRNA generates an exon3-skip mutant of PRDM1 in CAR-T, which leads to increased proliferation, stem-like properties, central memory and longevity in these cells, resulting in their high efficacy in vivo across multiple types of CAR-Ts and corresponding cancer models, including solid tumor. By mechanistic studies, we found that this Dexon3mutant removes a piece of PR domain of PRDM1. Via histone arrays, proteomics, epigenomics and transcriptomics, we found that PR domain interacts with histone H4 (HH4), and deletion of PR domain by Dexon3abolishes this interaction, leading to unleash of multiple transcription programs of multiple repressed T cell function genes particularly memory factors (direct targets of PRDM1). This set of studies unveiled previously unknown insights of T cell gene regulation, which can be harnessed to enhance T cell based cancer therapy.
Application of mass knock-in engineering can be broad and far-reaching. CLASH is not limited to encoding crRNA library; it can also mass-knock-in other pools such as GOF constructs, CAR pools, costim pools, and other elements. The modularity of these systems enables flexible and efficient generation of multiple different CARs in the same T cell, opening new capabilities of therapeutic cellular engineering with simplicity and precision, paving the way to engineer next-generation high-performance cell therapies to overcome the solid tumor challenge. The power of mass knock-in / CLASH can be further realized with in vivo settings. In Direct In-vivo CLASH setting, an RGN transgenic animal can be used. We have generated new RGN transgenic lines such as conditional and constitutive LbCpf1/LbCas12a and enAsCas12a mice mice (Tang et al. 2025 Nature Biomedical Engineering). Like Cas9 transgenic mice (Platt*, Chen* et al. 2014 Cell), we showed that Cpf1 mice simplifies multiplexed gene editing, accelerating immune cell gene editing and facilitates screening approaches. These mice can enable the powerful direct in vivo targeting research.
Publications
Dai X*, Park JJ*, Du Y*, Na Z*, Lam SZ*, Chow RD, Gu J, Renauer PA, Xin S, Chu Z, Liao C, Clark P, Zhao H, Slavoff S and Chen S†.
Massively parallel knock-in engineering of human T cells.
Nature Biotechnology, 2023 Jan 26. doi: 10.1038/s41587-022-01639-x.PMID: 36702900
https://www.nature.com/articles/s41587-022-01639-x
Kaiyuan Tang, Liqun Zhou, Xiaolong Tian, Shao-Yu Fang, Erica Vandenbulcke, Andrew Du, Johanna Shen, Hanbing Cao, Jerry Zhou, Krista Chen, Hyunu R. Kim, Zhicheng Luo, Shan Xin, Shawn H. Lin, Daniel Park, Luojia Yang, Yueqi Zhang, Kazushi Suzuki, Medha Majety, Xinyu Ling, Stanley Z. Lam, Ryan D. Chow, Ping Ren, Bo Tao, Keyi Li, Adan Codina, Xiaoyun Dai, Xingbo Shang, Suxia Bai, Timothy Nottoli, Andre Levchenko, Carmen J. Booth, Chen Liu, Rong Fan, Matthew B. Dong†, Xiaoyu Zhou† & Sidi Chen†. Cas12a-knock-in mice for multiplexed genome editing, disease modelling and immune-cell engineering.
Nature Biomedical Engineering, 2025
https://www.nature.com/articles/s41551-025-01371-2.
Dai X*, Park JJ*, Du Y, Kim RK, Wang G, Errami Y and Chen S†.
One-step generation of modular CAR-T with AAV-Cpf1.
Nature Methods (2019) Mar;16(3):247-254. doi: 10.1038/s41592-019-0329-7. PMID: 30804551.
https://www.nature.com/articles/s41592-019-0329-7
Program 3
Genome editing as new forms of immunotherapy modalities
This series of work innovates the concept of immune gene therapy by leveraging gene editing machinery directly as immunotherapy.
Cancer is a collection of complex diseases, each with an interwoven set of pathways that fuels the growth, proliferation, invasion, metastasis, and maintenance of immunosuppressive tumor microenvironment (TME). In the past decade, immunotherapy has transformed cancer treatment, turning several previously lethal cancers into manageable diseases for a subset of patients. Major types of immunotherapy include immune checkpoint blockade (ICB), cell therapy such as CAR-T, recombinant cytokines, oncolytic virus, and cancer vaccines. However, all current immunotherapy modalities face various limitations. Most of these patients do not respond to available therapies even in combinations2. Thus, there is a globally unmet clinical need of ultimate significance to develop new, better and safer immunotherapies for cancer patients.
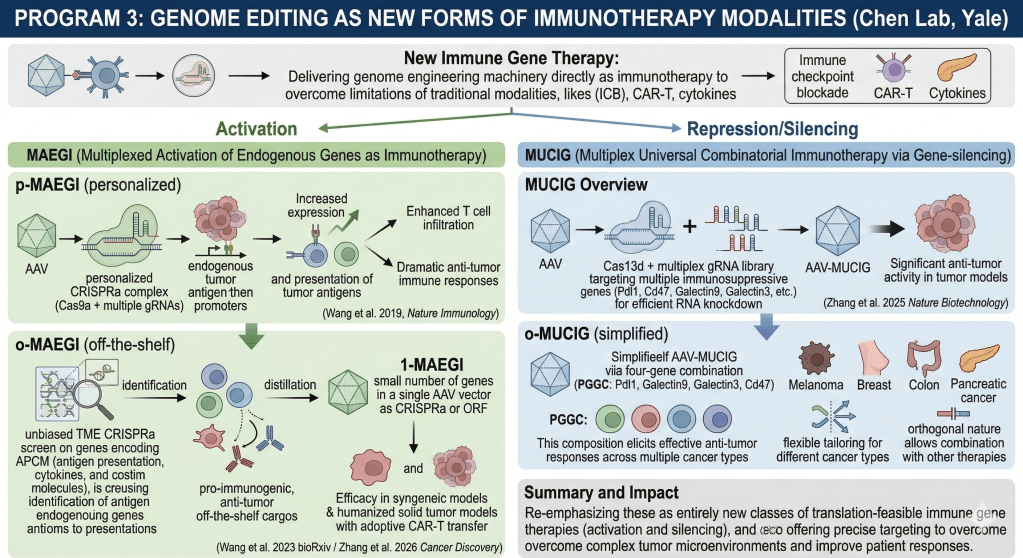
This urges for out-of-the-box thinking and development of an entirely new class of immunotherapy. Five years ago, I coined the concept of “immune gene therapy” by harnessing gene therapy as cancer immunotherapy. My team developed MAEGI (Multiplexed Activation of Endogenous Genes as Immunotherapy) as the first immune gene therapy that delivers genome engineering machinery itself as immunotherapy (Wang et al. 2019, Nature Immunology). MAEGI is a new form of immunotherapy that elicits anti-tumor immunity through multiplexed activation of endogenous genes. In personalized MAEGI (p-MAEGI), we leveraged CRISPR activation (CRISPRa) to directly augment the in situ expression of endogenous genes (thereby increasing the presentation of tumor antigens), leading to dramatic anti-tumor immune responses. This treatment modality led to alterations of the tumor microenvironment, marked by enhanced T cell infiltration and anti-tumor immune signatures.
In order to create off-the-shelf therapeutics and to simplify the manufacturing process, we recently advanced these concepts and developed off-the-shelf versions of MAEGI (o-MAEGI) that are more straightforward for translation (Wang et al. 2023 bioRxiv / Zhang et al. 2026 Cancer Discovery). The selection of fixed composition cargos for gene therapy is often by literature rationale and often only single genes. However, the number of immune genes that can be delivered or activated as potential therapeutics is large, and the combinations can increase exponentially. By devising an unbiased TME CRISPRa screen on genes encoding antigen presentation, cytokines and costim molecules (APCM), we have identified the o-MAEGI cargos that are pro-immunogenic and anti-tumor (Wang et al. 2023). We have been performing extensive optimization of the composition and further distilled to a small number of genes that can fit in one single AAV vector (1-MAEGI) for delivery as CRISPRa or simple ORFs. Our o-MAEGI and 1-MAEGI candidates showed significant efficacy in syngeneic models, as well as humanized solid tumor model with adoptive CAR-T cell transfer. o-MAEGI thus provide promising and translation-feasible candidates for next-gen immune gene therapy.
Suppressive TME encode multiple immunosuppressive factors including immune checkpoint proteins. To develop an approach that enable flexible silencing of desired combinations of such factors, we develop gene-silencing based immune gene therapy. We develop MUCIG (Multiplex Universal Combinatorial Immunotherapy via Gene-silencing) as a new form of versatile cancer immunotherapy (Zhang et al. 2025 Nature Biotechnology). We use the RNA knockdown capabilities of Cas13d to efficiently target multiple immunosuppressive genes on demand, to silence various combinations of multiple immunosuppressive factors in the TME. AAV-mediated administration of MUCIG (AAV-MUCIG) has significant anti-tumor activity with several compositions of Cas13d gRNA libraries. Further optimization results in a simplified off-the-shelf AAV-MUCIG with a four gene combination (PGGC: Pdl1, Galectin9, Galectin3 and Cd47) show that it can elicit effective anti-tumor responses across different tumor models of multiple cancer types, including melanoma, breast cancer, colon cancer, and pancreatic cancer. MUCIG allows flexible tailoring of immune gene silencing combinations for different cancer types, different sub-types, and patient populations for more precise targeting. Because this is an orthogonal (gene silencing as immunotherapy), MUCIG can be used in combination with virtually any other therapeutic modalities.
Publications
Wang G*, Chow RD*, Bai Z, Zhu L, Errami Y, Dai X, Dong MB, Ye L, Zhang X, Renauer RA, Park JJ, Shen L, Ye H, Fuchs CS, and Chen S†.
Multiplexed activation of endogenous genes by CRISPRa elicits potent anti-tumor immunity.
Nature Immunology 20, 1494–1505 (2019). published online14 October 2019 DOI: https://doi.org/10.1038/s41590-019-0500-4
https://www.nature.com/articles/s41590-019-0500-4
Guangchuan Wang, Feifei Zhang, Ryan D Chow, Emily He, Lvyun Zhu, Qin Han, Sidi Chen †
Rational design of immune gene therapy combinations via in vivo CRISPR activation screen of tumor microenvironment modulators
bioRxiv 2023
doi: https://doi.org/10.1101/2023.03.14.532665
https://www.biorxiv.org/content/10.1101/2023.03.14.532665v1
Feifei Zhang, Guangchuan Wang, Ryan D Chow, Emily He, Medha Majety, Yueqi Zhang, Sidi Chen †
Multiplexed repression of immunosuppressive genes as combinatorial cancer immunotherapy
bioRxiv 2023
doi: https://doi.org/10.1101/2023.03.14.532668
https://www.biorxiv.org/content/10.1101/2023.03.14.532668v1
Feifei Zhang, Ryan Chow, Emily He, Chuanpeng Dong, Shan Xin, Daniyal Mirza, Yanzhi Feng, Xiaolong Tian, Nipun Verma, Medha Majety, Yueqi Zhang, Guangchuan Wang†, Sidi Chen†. Multiplexed inhibition of immunosuppressive genes with Cas13d for combinatorial cancer immunotherapy.
Nature Biotechnology, 2025.
https://www.nature.com/articles/s41587-024-02535-2.
Program 4
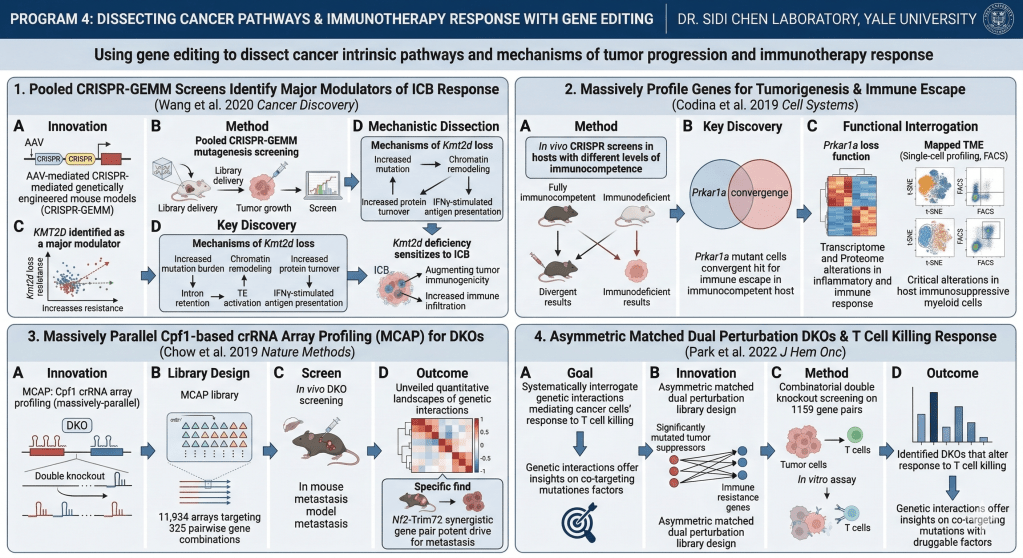
Using gene editing to dissect cancer intrinsic pathways and mechanisms of tumor progression and immunotherapy response
This series of work leverages unbiased genetic screen to identify critical genes controlling tumor progression and immunotherapy response, and uses gene editing to dissect cancer intrinsic pathways and mechanisms.
Immune checkpoint blockade (ICB) has shown remarkable clinical efficacy. However, only a fraction of patients respond to ICB. We invented pooled CRISPR-mediated genetically engineered mouse models (CRISPR-GEMM) models that allow direct in vivo dissection of causal genes of oncogenesis and tumor progression (Chow et al. 2017 Nature Neuroscience; Wang et al. 2018 Science Advances). We performed CRISPR-GEMM screen to identify factors governing sensitivity and resistance to ICB (Wang et al. 2020 Cancer Discovery), and identified KMT2D as a major modulator of ICB response. Our mechanistic dissection revealed that Kmt2d loss led to increased DNA damage and mutation burden, chromatin remodeling, intron retention, activation of transposable elements, increased protein turnover and IFNγ-stimulated antigen presentation. Kmt2d-mutant tumors have increased immune infiltration across multiple cancer types. Kmt2d deficiency sensitizes tumors to ICB by augmenting tumor immunogenicity.
Using in vivo CRISPR screens, we massively profiled genes that drive tumorigenesis and immune escape in hosts with different levels of immunocompetence (Codina et al. 2019 Cell Systems). As a convergent hit, Prkar1a mutant cells escape immune checks in fully immunocompetent hosts. Functional interrogation showed that Prkar1a loss altered the transcriptome and proteome in inflammatory and immune responses and extracellular proteins. Single-cell profiling and FACS mapped the TME of Prkar1a mutant tumors and revealed critical alterations in host immunosuppressive myeloid cells.
To study complex genetic interactions, we scaled up the dimension of CRISPR screens. We developed massively-parallel Cpf1/Cas12a crRNA array profiling (MCAP) for interrogation of double knockouts (DKOs) (Chow et al. 2019 Nature Methods). We designed an MCAP library of 11,934 arrays targeting 325 pairwise gene combinations and performed in vivo double knockout screening. We unveiled quantitative landscapes of genetic interactions and validated Nf2-Trim72 as a potent synergistic gene pair that drive metastasis. To systematically interrogate genetic interactions mediating cancer cells’ response to T cell killing, we designed an asymmetric dual perturbation library targeting the matched combinations between significantly mutated tumor suppressors and immune resistance genes (Park et al. 203 Journal of Hematology and Oncology). We performed combinatorial double knockout screening on 1159 gene pairs and identified DKOs that altered response to T cell cytotoxicity. Such genetic interactions offer insights on co-targeting clinically relevant mutations with causal druggable factors.
Publications
Wang G*, Chow RD*, Zhu L*, Bai Z*, Ye L, Zhang F, Renauer RA, Dong MB, Dai X, Zhang X, Du Y, Cheng Y, Niu L, Chu Z, Kim K, Liao C, Clark P, Errami Y and Chen S†.
CRISPR-GEMM pooled mutagenic screening identifies KMT2D as a major modulator of immune checkpoint blockade.
Cancer Discovery (2020) published on September 4, 2020; doi: 10.1158/2159-8290.CD-19-1448
Park JJ*, Codina A*, Ye L*, Lam SZ, Guo J, Clark P, Zhou X, Peng L, Chen S†.
Double knockout CRISPR screen in cancer resistance to T cell cytotoxicity.
Journal of Hematology & Oncology volume 15, Article number: 172 (2022) Dec 1;15(1):172. doi: 10.1186/s13045-022-01389-y. PMID: 36456981
** Initial bioRxiv deposit
Double knockout CRISPR screen in cancer resistance to T cell cytotoxicity
Jonathan J. Park, Adan Codina, Lupeng Ye, Stanley Lam, Jianjian Guo, Paul Clark, Xiaoyu Zhou, Lei Peng, Sidi Chen
doi: https://doi.org/10.1101/2022.03.01.482556
https://www.biorxiv.org/content/10.1101/2022.03.01.482556v1
Chow RD*, Wang G*, Ye L*, Codina A, Kim RK, Shen L, Dong M, Errami Y and Chen S†.
High-density in vivo profiling of metastatic double knockouts through CRISPR-Cpf1.
Nature Methods (2019) May;16(5):405-408. doi: 10.1038/s41592-019-0371-5. Epub 2019 Apr 8.
** Initial bioRxiv deposit
Mapping in vivo genetic interactomics through Cpf1 crRNA array screening
Ryan D. Chow, Guangchuan Wang, Adan Codina, Lupeng Ye, Sidi Chen
doi: https://doi.org/10.1101/153486
https://www.biorxiv.org/content/10.1101/153486v1.full
Codina A*, Renauer P*, Wang G*, Chow RD*, Park JJ, Ye H, Zhang K, Dong M, Gassaway B, Ye L, Errami Y, Shen L, Chang A, Jain D, Herbst RS, Bosenberg M, Rinehart J, Fan R and Chen S†.
Convergent identification and interrogation of tumor-intrinsic factors that modulate cancer immunity in vivo. Cell Systems (2019) Feb 27;8(2):136-151.e7. doi: 10.1016/j.cels.2019.01.004. Epub 2019 Feb 20. PMID: 30797773; PMCID: PMC6592847.
https://linkinghub.elsevier.com/retrieve/pii/S2405-4712(19)30032-8
Xue W*, Chen S*, Yin H*, Tammela T, Papagiannakopoulos T, Joshi NS, Cai W, Yang G, Bronson R, Crowley DG, Zhang F, Anderson DG, Sharp PA, Jacks T. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature. 2014 Oct 16;514(7522):380-4. doi: 10.1038/nature13589. PMID: 25119044.
https://pubmed.ncbi.nlm.nih.gov/25119044/
Platt RJ*, Chen S*, Zhou Y, Yim MJ, Swiech L, Kempton HR, Dahlman JE, Parnas O, Eisenhaure TM, Jovanovic M, Graham DB, Jhunjhunwala S, Heidenreich M, Xavier RJ, Langer R, Anderson DG, Hacohen N, Regev A, Feng G, Sharp PA, Zhang F. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell. 2014 Oct 9;159(2):440-55. doi: 10.1016/j.cell.2014.09.014. PMID: 25263330 (Cover story).
https://www.cell.com/fulltext/S0092-8674(14)01163-5
Chow RD*, Guzman CD*, Wang G*, Schmidt F*, Youngblood MW, Ye L, Errami Y, Dong MB, Martinez MA, Zhang S, Renauer P, Bilguvar K, Gunel M, Sharp PA, Zhang F, Platt RJ †, Chen S †. AAV-mediated direct in vivo CRISPR screen identifies functional suppressors in glioblastoma. Nature Neuroscience, 20, 1329–1341 (2017) doi:10.1038/nn.4620)
https://www.nature.com/articles/nn.4620
Wang G*, Chow RD*, Ye L, Guzman CD, Dai X, Dong MB, Zhang F, Sharp PA, Platt RJ†, and Chen S†. Mapping a Functional Cancer Genome Atlas of Tumor Suppressors in Mouse Liver Using AAV-CRISPR Mediated Direct in vivo Screening. (2018) Science Advances. Feb 28;4(2):eaao5508. PMID: 29503867.
https://www.science.org/doi/10.1126/sciadv.aao5508
Chen S*, Sanjana NE*, Zheng K, Shalem O, Lee K, Shi X, Scott DA, Song J, Pan JQ, Weissleder R, Lee H, Zhang F, Sharp PA. Genome-wide CRISPR screen in a mouse model of tumor growth and metastasis. Cell. 2015 Mar 12;160(6):1246-60. PMID: 25748654; (* = co-first authors) (Best of Cell2015)
https://www.cell.com/fulltext/S0092-8674(15)00204-4
Chow RD*, Wang G*, Ye L*, Codina A, Kim RK, Shen L, Dong M, Errami Y and Chen S†. High-density in vivo profiling of metastatic double knockouts through CRISPR-Cpf1. Nature Methods (2019) May;16(5):405-408. doi: 10.1038/s41592-019-0371-5.
https://www.nature.com/articles/s41592-019-0371-5
Chow RD and Chen S†. Sno-derived RNAs are prevalent molecular markers of cancer immunity. Oncogene, 2018 DOI – 10.1038/s41388-018-0420-z, PMID: 30072739
https://www.nature.com/articles/s41388-018-0420-z
Chow RD and Chen S†. Cancer CRISPR screens in vivo. Trends In Cancer. 2018 May;4(5):349-358. doi: 10.1016/j.trecan.2018.03.002. Review. PMID: 29709259 (Cover story)
https://www.sciencedirect.com/science/article/abs/pii/S2405803318300591
Chow RD*, Chen JS*, Shen J, and Chen S†. A web tool for the design of prime-editing guide RNAs. (pegFinder paper). Nature Biomedical Engineering (2020) DOI: https://doi.org/10.1038/s41551-020-00622-8
https://www.nature.com/articles/s41551-020-00622-8
Program 5
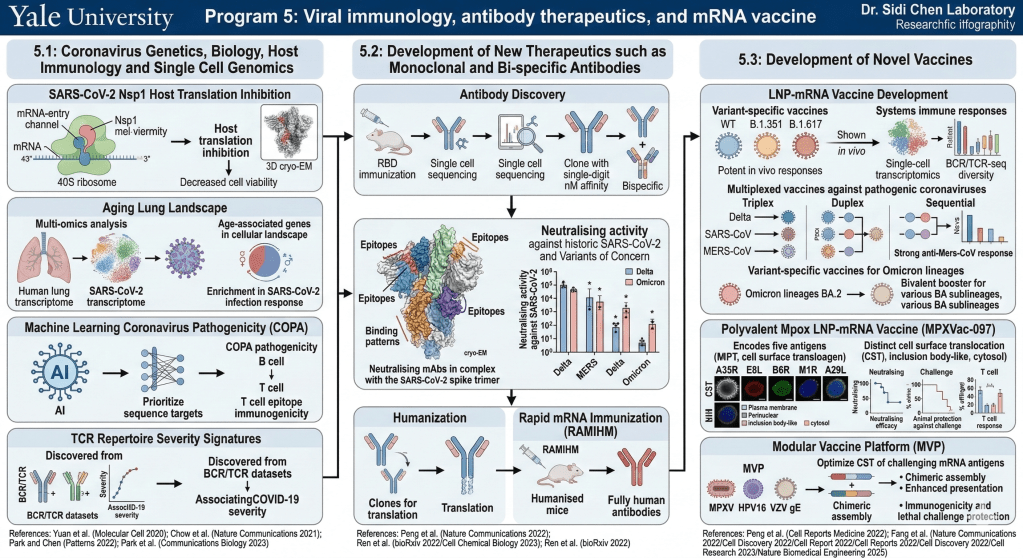
Viral immunology, antibody therapeutics, and mRNA vaccine
5.1. Coronavirus genetics, biology, host immunology and single cell genomics
We perform genomic analysis of expression and immune signatures to identify genetic links to disease vulnerability factors such as age; and understanding of viral immunology for the development of better coronavirus vaccines.The causative virus of the COVID-19 pandemic, SARS-CoV-2, uses its nonstructural protein 1 (Nsp1) to suppress cellular, but not viral, protein synthesis through yet unknown mechanisms. We show here that among all viral proteins, Nsp1 has the largest impact on host viability in the cells of human lung origin. Differential expression analysis of mRNA-seq data revealed that Nsp1 broadly alters the cellular transcriptome. Our cryo-EM structure of the Nsp1-40S ribosome complex shows that Nsp1 inhibits translation by plugging the mRNA-entry channel of the 40S. Our results elucidate the mechanism of host translation inhibition by SARS-CoV-2 and advances the understanding of the impacts from a major pathogenicity factors of SARS-CoV-2.Our integrative multi-omics analysis also mapped out the aging transcriptome and cellular landscape of the human lung in relation to SARS-CoV-2. Surprisingly, age-associated genes are heavily enriched among those induced or suppressed by SARS-CoV-2 infection. We recently developed a set of machine learning approaches that distinguishes pathogenic sequence features of coronaviruses down to base pair and amino acid resolutions with an integrated coronavirus pathogenicity (COPA) score. We identified pathogenic hotspots across the SARS-CoV-2 viral genome including a large number of previously unappreciated features in viral proteins. Our integrated pathogenicity-linked genomic profiles with B cell and T cell epitope immunogenicity features for prioritization and enrichment of key sequence targets that can guide vaccine development (Park and Chen, 2022, Patterns). We also analyzed T cell receptor repertoires of large single cell and BCR/TCR datasets of COVID-19 patients and healthy donors, and discovered signatures associated with COVID-19 severity.
Yuan S*, Peng L*, Park JJ, Hu Y, Devarkar SC, Dong MB, Wu S, Chen S†, Lomakin I† and Xiong Y†.
Nonstructural protein 1 of SARS-CoV-2 is a potent pathogenicity factor redirecting host protein synthesis machinery toward viral RNA. Molecular Cell. (2020) DOI: https://doi.org/10.1016/j.molcel.2020.10.034
https://www.sciencedirect.com/science/article/pii/S1097276520307413
Chow RD and Chen S†.
The aging transcriptome and cellular landscape of the human lung in relation to SARS-CoV-2. doi: 10.1101/2020.04.07.030684. Nature Communications (2021)
https://www.nature.com/articles/s41467-020-20323-9
Park JJ and Chen S†. Metaviromic identification of genetic hotspots of coronavirus pathogenicity using machine learning. Patterns 2022. doi:10.1016/j.patter.2021.100407.
Park JJ, Lee KAV, Lam SZ, Chen S†. Machine learning identifies T cell receptor repertoire signatures associated with COVID-19 severity. Communications Biology 2023.
https://www.nature.com/articles/s42003-023-04447-4
5.2. Development of new therapeutics such as monoclonal and bi-specific antibodies
Our laboratory is conducting development of novel therapeutic candidates such as neutralizing antibodies. Effective therapeutic options such as monoclonal antibodies (mAbs) are still critical. Here, we report the development, cryo-EM structures, and functional analyses of mAbs that potently neutralize SARS-CoV-2 variants of concern. By high-throughput single cell sequencing of B cells from spike receptor binding domain (RBD) immunized animals, we identified two highly potent SARS-CoV-2 neutralizing mAb clones that have single-digit nanomolar affinity and low-picomolar avidity, and generated a bispecific antibody. Lead antibodies showed strong inhibitory activity against historical SARS-CoV-2 and several emerging variants of concern. We solved several cryo-EM structures at ∼3 Å resolution of these neutralizing antibodies in complex with prefusion spike trimer ectodomain, and revealed distinct epitopes, binding patterns, and conformations. The lead clones also showed potent efficacy in vivo against authentic SARS-CoV-2 in both prophylactic and therapeutic settings. We also generated and characterized a humanized antibody to facilitate translation and drug development. The humanized clone also has strong potency against both the original virus and the B.1.617.2 Delta variant. We also recently developed fully human / humanized monoclonal and bispecific mAbs against the Omicron lineages. These mAbs expand the repertoire of therapeutics against SARS-CoV-2 and emerging variants.
Peng L*, Hu Y*, Mankowski MC*, Ren P*, Chen RE, Wei J, Zhan M, Li T, Tripler T, Ye L, Chow RD, Fang Z, Wu C, Dong MB, Cook M, Wang G, Clark P, Nelson B, Klein D, Sutton R, Diamond MS, Wilen CB†, Xiong Y†, Chen S†. Monospecific and bispecific monoclonal SARS-CoV-2 neutralizing antibodies that maintain potency against B.1.617. Nature Communications 2022
https://www.nature.com/articles/s41467-022-29288-3
Ren P*, Peng L*, Fang Z, Suzuki K, Renauer PA, Lin Q, Bai M, Yang L, Li T, Clark P, Klein D, Chen S†. First published as a preprint:Potent and specific human monoclonal antibodies against SARS-CoV-2 Omicron variant by rapid mRNA immunization of humanized mice. bioRxiv 2022.03.17.484817; doi: https://doi.org/10.1101/2022.03.17.484817 RAMIHM generates fully human monoclonal antibodies by rapid mRNA immunization of humanized mice and BCR-seq. Cell Chemical Biology 2023. Cover Article.
https://www.sciencedirect.com/science/article/pii/S245194562200455X
Ren P, Hu Y, Peng L, Yang L, Suzuki K, Fang Z, Bai M, Zhou L, Feng Y, Zou Y, Xiong Y†, Chen S†. Function and Cryo-EM structures of broadly potent bispecific antibodies against multiple SARS-CoV-2 Omicron sublineages. bioRxiv 2022 PMID: 35982661. doi: https://doi.org/10.1101/2022.08.09.503414
5.3. Development of novel vaccines
Lipid-nanoparticle(LNP)-mRNA vaccines offer protection against COVID-19. However, multiple variant lineages caused widespread breakthrough infections. To generate and test variant-specific vaccines, we generated LNP-mRNAs specifically encoding wildtype, B.1.351 and B.1.617 SARS-CoV-2 spikes, and systematically studied their immune responses in animal models. All three LNP-mRNAs induced potent antibody responses in mice. However, WT-LNP-mRNA vaccination showed reduced neutralization against B.1.351 and B.1.617; and B.1.617-specific vaccination showed differential neutralization. All three vaccine candidates elicited antigen-specific CD8 and CD4 T cell responses. Single cell transcriptomics of B.1.351-LNP-mRNA and B.1.617-LNP-mRNA vaccinated animals revealed a systematic landscape of immune cell populations and global gene expression. Variant-specific vaccination induced a systemic increase in reactive CD8 T cell population, with a strong signature of transcriptional and translational machineries in lymphocytes. BCR-seq and TCR-seq unveiled repertoire diversity and clonal expansions in vaccinated animals. These data provide direct systems immune profiling of variant-specific LNP-mRNA vaccination in vivo. Although successful COVID-19 vaccines have been developed, multiple pathogenic coronavirus species exist, urging for development of multi-species coronavirus vaccines. We developed prototype LNP-mRNA vaccine candidates against SARS-CoV-2 (Delta variant), SARS-CoV and MERS-CoV, and test how multiplexing of these LNP-mRNAs can induce effective immune responses in animal models. A triplex scheme of LNP-mRNA vaccination induced antigen-specific antibody responses against SARS-CoV-2, SARS-CoV and MERS-CoV, with a relatively weaker MERS-CoV response in this setting. Duplex LNP-mRNA vaccination showed potent anti-MERS-CoV immune responses, without sacrificing the effect of companion SARS-CoV-2 or SARS-CoV vaccination in the combination. Sequential vaccination showed potent antibody responses against all three species, significantly stronger than simultaneous vaccination in mixture. Single cell RNA-seq profiled the global systemic immune repertoires and the respective transcriptome signatures of multiplexed vaccinated animals, which revealed a systemic increase in activated B cells, and differential gene expression signatures across major adaptive immune cells.
Peng, L.*, Renauer, P.A.*, Ökten, A.*, Fang, Z.*, Park, J.J.*, Zhou, X.*, Lin, Q., Dong, M.B., Filler, R., Xiong, Q., Clark, P., Lin, C., Wilen, C.B. †, Chen, S. †. Variant-specific vaccination induces systems immune responses and potent in vivo protection against SARS-CoV-2. Cell Reports Medicine (2022), doi: https://doi.org/10.1016/j.xcrm.2022.100634.
Fang Z*, Peng L*, Filler R, Suzuki K, McNamara A, Lin Q, Renauer PA, Yang L, Menasche M, Sanchez A, Ren P, XionQg, Strine M, Clark P, Lin C, Ko AI, Grubaugh NG, Wilen CB, and Chen, S. †. Omicron-specific mRNA vaccination alone and as a heterologous booster against SARS-CoV-2. doi: https://doi.org/10.1101/2022.02.14.480449 | Nature Communications (2022)
https://www.nature.com/articles/s41467-022-30878-4
Fang Z*, Peng L*, Lin Q, Zhou L, Yang L, Feng Y, Ren P, Renauer PA, Park JJ, Zhou X, Wilen CB, Chen S†. Heterotypic vaccination responses against SARS-CoV-2 Omicron BA.2. doi: https://doi.org/10.1101/2022.03.22.485418 Cell Discovery (2022)
https://www.nature.com/articles/s41421-022-00435-w
Peng L*, Fang Z*, Renauer PA*, McNamara A, Park JJ, Lin Q, Zhou X, Dong MB, Zhu B, Zhao H, Wilen CB, Chen S. † Multiplexed LNP-mRNA vaccination against pathogenic coronavirus species.doi: https://doi.org/10.1101/2022.05.07.491038 Cell Reports (2022)
Fang Z, Monteiro VS, Hahn AM, Grubaugh ND, Lucas C, Chen S. Bivalent mRNA vaccine booster induces robust antibody immunity against Omicron lineages BA.2, BA.2.12.1, BA.2.75 and BA.5. Cell Discovery. 2022 Oct 11; 8: 108
https://www.nature.com/articles/s41421-022-00473-4
The global monkeypox (Mpox) outbreak leads to a surge in demand for vaccine. While mRNA vaccines are successful against SARS-CoV-2, their efficacy against other emerging pathogens, such as the monkeypox virus (MPXV), remains unexplored. We developed a polyvalent mRNA lipid nanoparticle (LNP) vaccine candidate (MPXVac-097) that encodes five MPXV antigens and assess the immunogenicity in animal models. Confocal imaging and flow cytometry show that all five MPXV antigens encoded by MPXVac-097 are stably expressed, with A35R and E8L enriched in the plasma membrane, B6R in perinuclear structures, M1R in plasma membrane or inclusion body-like structures, and A29L in the cytosol. Upon immunization, MPXVac-097 vaccine elicits variable antibody response to the five MPXV antigens, yet has efficient neutralization against MPXV and other poxviruses, and protects animals against challenge against vaccinia virus. Bulk T cell receptor (TCR) sequencing of peripheral T cells from MPXVac-097 vaccinated mice reveals preferential usage of VJ sequences and clonal expansion. Mice vaccinated with MPXVac-097 show no significant pathology, and little changes in blood cell counts and liver enzymes compared to untreated. Together, these data provide the efficacy, immunogenicity and preliminary safety of MPXVac-097 to support its developability as an Mpox mRNA vaccine.
Fang Z*, Monteiro VS*, Renauer PA, Shang X, Suzuki K, Vogels C, Ling X, Bai M, Xiang Y, Levchenko A, Booth CJ, Lucas C†, and Chen S†. Polyvalent mRNA vaccination elicited potent immune response to monkeypox virus surface antigens. Cell Research (2023) 0:1–4; https://doi.org/10.1038/s41422-023-00792-5
Certain messenger RNA antigens in mRNA vaccines elicit an insufficient immune response due to challenges in cell surface translocation (CST) of the antigens. Here we develop a modular vaccine platform (MVP) to enhance the immunogenicity of challenging mRNA antigens by optimizing antigen expression and presentation. MVPs enable the modular assembly of chimeric antigens. Our platform comprises diverse modules capable of generating >2,500 combinations with any antigen and displaying distinct antigen epitopes on the cell surface. We quantify the CST efficacy of various modules using multiple antigens, including the mpox virus (MPXV) proteins A29, M1R and A35R, and compare chimeric antigen surface expression in multiple cell lines. Using MPXV as a model, we identify optimal modules that enhance the CST of multiple MPXV antigens, improving the immune response of lipid nanoparticle mRNAs and protecting against lethal viral challenge. With these effective CST modules, we further demonstrate the generalizability of MVP by optimizing additional mRNA antigens, including the human papillomavirus 16 proteins E6 and E7 and the varicella zoster virus glycoprotein gE. This platform is applicable to any antigen of interest, facilitating the development of mRNA vaccines against challenging targets.
Zhenhao Fang, Valter S. Monteiro, Changin Oh, Kawthar Al Janabi, Luciano Romero, Nabihah Ahsan, Luojia Yang, Lei Peng, Daniel DiMaio, Carolina Lucas†, Sidi Chen†. A modular vaccine platform for optimized lipid nanoparticle mRNA immunogenicity.
Nature Biomedical Engineering, 2025.